Our Mission
The GarMiglio Lab’s mission is deciphering the molecular mechanisms of cancer progression, metastasis, and resistance to therapy in both adult and pediatric brain cancer as well as in other solid tumors including lung, breast and melanoma.
We combine computational approaches and experimental models to:
- Dissect the complexity of the tumor ecosystem by identifying functionally relevant cell subpopulations and decipher their molecular traits, functions, spatial organization and cell-cell interaction through single cell and spatial technologies.
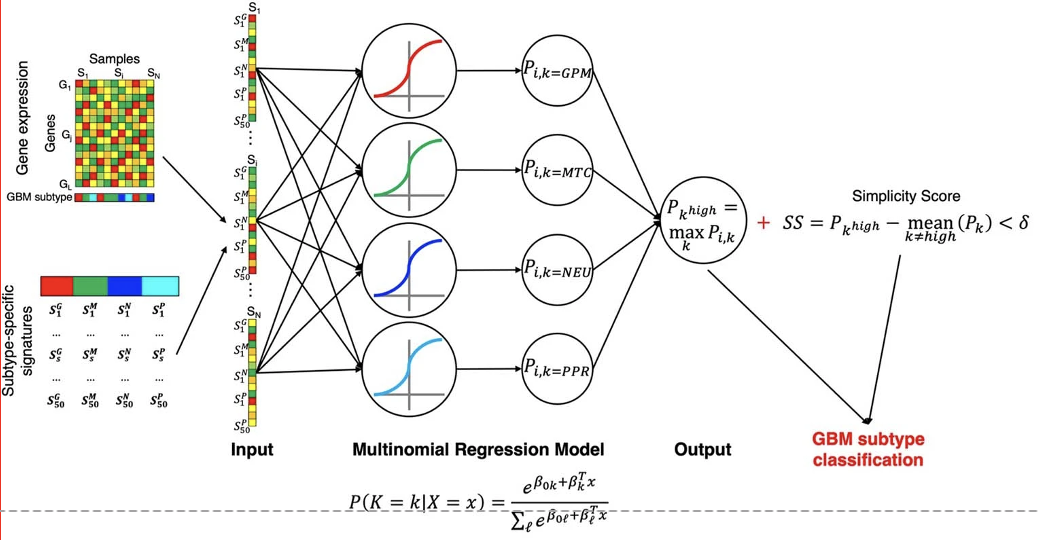
- Define clinically relevant subtypes of tumors by integrating bulk-level multi-omics including transcriptomics, epigenetics, genomics, phospho-proteomics profiles from hundreds to thousands tumors using machine learning approaches and reconstruction of gene/protein regulatory networks.
- Implement experimental models to recapitulate the human tumor ecosystem to translate our findings into real-world applications and develop novel therapeutic strategies for cancer patients.
"Our team embraces a collaborative approach, working closely with researchers and clinicians. Our final aim is to develop more effective therapies and improve the survival outcome of patients including children affected by this deadly disease."
Our Team
Led by principal investigators dedicated to precision oncology and computational biology.

Luciano Garofano, PhD
Principal Investigator
Luciano Garofano is an Assistant Professor at The Translational Genomics Research Institute (TGen). He earned his Ph.D. in Bioinformatics in 2017 at the University of Sannio, Italy. His PhD focused on the development of machine learning approaches for the reconstruction of gene- and protein-regulatory networks to unravel the heterogenous biological and genetic mechanisms underlying distinct subgroups of solid tumors.
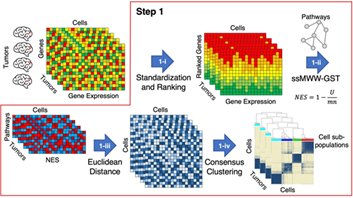
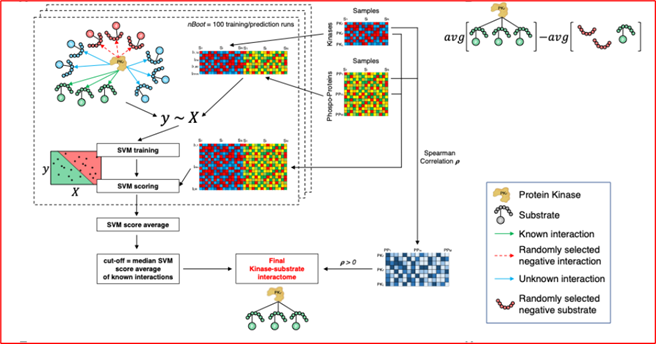
During his post-doctoral journey mentored by Dr. Antonio Iavarone, Dr. Garofano has designed several computational tools, including RGBM (Regularized Gradient Boosting Machine), scBiPaD (single-cell Biological Pathway Deconvolution) and SPHINKs (Substrate PHosphosite-based Inference for Network of Kinases) and informed key therapeutic vulnerabilities and functionally-relevant features of adult and pediatric brain tumors.

Simona Migliozzi, PhD
Principal Investigator
Assistant Professor at The Translational Genomics Research Institute (TGen). Native of Italy, she completed her Ph.D. in Molecular Oncology at University of Catanzaro (2019). Her dissertation work focused on the development of computational approaches to integrate NGS data for the molecular characterization of different subtypes of solid tumors (glioma, ovarian, colon, breast cancers).
In 2019, she joined Dr. Antonio Iavarone’s laboratory at Columbia University and University of Miami as a postdoctoral fellow. Her work focused on developing machine learning approaches using single cell and multi-omics data to dissect heterogeneity of glioblastoma multiformeand identify clinically-relevant subtypes.
Research Projects
Exploring the frontiers of cancer biology through specific lines of inquiry.
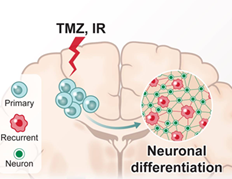
Dissecting glioma ecosystem during evolution
Our previous studies have shown that glioma ecosystem is dramatically changing during evolution. We want seek the biological mechanisms driving changes in tumor composition, tumor microenvironment and their spatial patterns to identify new therapeutic vulnerabilities.
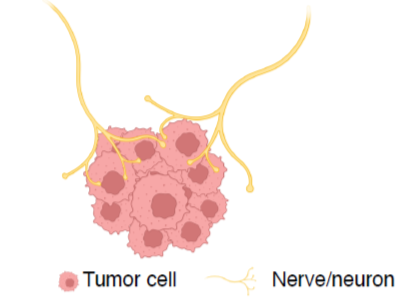
Role of neurons-cancer cross-talk in non-CNS tumor progression
We showed the existence of a tumor neuronal-like program in lung and breast cancer. We test whether the innervation of non-CNS tumors (gastrointestinal, pancreatic, prostate) by peripheral nerves regulates cancer progression, seeking molecular mechanisms driving neuron-cancer interactions.
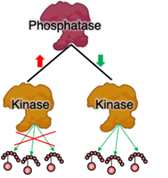
Reconstructing spatiotemporal proteomic mosaicism in pediatric brain tumors
We have shown an evolutionary trajectory of distinct tumor cellular states and TME cell subpopulations from low to high-grade pediatric tumors. We seek to identify critical post-translational regulators (kinases, ubiquitin ligases) responsible for the biological make-up of children's brain tumors.
Software & Tools
Open source computational tools developed by our lab for the scientific community.
Selected Publications
Key research findings and scientific contributions.

Parsing glioblastoma heterogeneity for redefined classification
Analysis of glioblastoma heterogeneity can classify the disease according to four fundamental functional properties, depicted here as branches of one tree. Genetic alterations common to these four subtypes are within the glioblastoma trunk, but each subtype diverges through distinct genetic lesions and gene-expression programs that incorporate prognostic and therapeutic attributes.
Garofano L.*, Migliozzi S.*, Oh*, et al., Nature Cancer (2021)
Link
Identifying master kinases in cancer with integrative multi-omics
A kinase–substrate–phosphosite network interactome identifies master kinases specific for glioblastoma subtypes, as potential therapeutic targets.
Migliozzi S.*, Oh*, Hasanain*, Garofano L.*, et al., Nature Cancer (2023)
Link
Restraint of cancer cell plasticity by spatial homotypic clustering
Among carnivorous plants, two wilted heads of daisies embody homotypic clusters of malignant cells, still holding a single identity despite decline. Around them, the open traps of insect-eating plants evoke dispersed agglomerates in which identity dissolves and invasive programs emerge. The juxtaposition of a fading but coherent mass with predatory heterogeneity symbolizes the finding of Migliozzi et al. that the spatial self organization of malignant cell subtypes is not neutral but can preserve or release the plastic potential through which glioblastoma advances.
Migliozzi S.*, Adabbo B.*, Garofano L.*, et al. Cancer Cell (2025)
LinkMore Publications
Kim K.H.*, Migliozzi S.*, […], Garofano L., […], et al. Integrated proteogenomic characterization of glioblastoma evolution. Cancer Cell (2024)
PaperGarofano L., D’Angelo F., Oh M., Ceccarelli M., Bielle F., Sanson M., Lasorella A, Iavarone A. Identification of distinct tumor-TME ecomodules in glioma from Neurofibromatosis type 1. Nature Medicine (in submission)
D’Angelo F., […], Garofano L., […], et al. The molecular landscape of glioma in patients with Neurofibromatosis 1. Nature Medicine (2019)
PaperNomura M.*, Spitzer A.*, Johnson K.C.*, Garofano L.*, Migliozzi S., […], Lasorella A., Verhaak R.G.W., Iavarone A., Suvà M.L., Tirosh I. The Multi-Layered Transcriptional Architecture of Glioblastoma Ecosystems. Nature Genetics (2025)
PaperJoin Our Team
We are looking for passionate, highly motivated and talented staff computational scientists or postdoctoral researchers. Candidates with PhD or Master’s Degree in statistics, mathematics, computational biology, bioinformatics and informatics with strong passion for cancer biology are welcome to contact us and potentially join our team. One to two years research lab experience are required.
Lab’s mission and values
Professional Development
Our role as PIs is to support your professional development through individualized training plan and help you reach your career goals. We will help you promoting your work, establishing a network, and be an advocate for larger administrative issues.
Regular Meetings
We will have regular weekly lab meetings to review the data generated, troubleshoot any technical problems, brainstorm together and establish research plans for the subsequent week. Everyone’s opinion is valuable.
Collaboration & Teamwork
Collaboration and teamwork are the keys to impactful research. We seek candidates who are willing to help others as well as work together with researchers with different expertise and clinicians to achieve scientific excellence.
Open Communication
Our lab has open door policy to encourage open communication. You can talk to us about any professional or personal issues.
"Our lab is committed to fostering a respectful, collaborative, and welcoming environment where individuals from all backgrounds can thrive and contribute meaningfully to scientific discovery."
Current Openings
Computational Scientist Roles
GarMiglio Lab
The GarMiglio Lab led by Dr. Luciano Garofano and Dr. Simona Migliozzi at The Translational Genomics Research Institute is an interdisciplinary research group aimed at understanding the molecular mechanisms of cancer progression, metastasis, and resistance to therapy in solid tumors, with particular focus on adult and pediatric brain tumors, lung, breast, melanoma, and brain metastasis.
Our groups combine patient biospecimens, computational approaches, and experimental models to dissect cancer heterogeneity, identify functional and clinically relevant tumor subtypes, and extract targetable molecular nodes, with the final aim of developing more effective therapies that improve survival outcomes for cancer patients.
Ongoing projects include:
- Dissecting glioma ecosystem during evolution
- Role of neurons-cancer crosstalk in non-CNS tumor progression
- Reconstructing the spatiotemporal proteomic mosaicism in pediatric brain tumors
We are looking for talented researchers to develop computational pipelines and implement analytical models to integrate multi-omics data. Candidates with expertise in machine learning approaches and reconstruction of gene/protein regulatory networks, and with knowledge of or strong interest in cancer biology, are encouraged to apply.
The successful candidate will take a leadership role in ongoing projects as well as develop new research ideas. Specific duties will include data analysis, interpretation of results, presentation of findings, and preparation of peer-reviewed manuscripts and grant proposals.
Required
- Master's or PhD in Computational Biology, Bioinformatics, Computer Science, Statistics, or related
- Deep proficiency or strong interest in molecular cancer biology
- Excellent communication and organizational skills
- Ability to manage time and multitask effectively
Preferred
- At least 1 year of research experience
- Experience coding in bash, Python and/or R
- Experience with Bioconductor packages
- Confidence operating cluster systems (SLURM/PBS)
Application instructions
Please apply by emailing a cover letter (Statement of Research, optional), your CV, and contact information for two references to: smigliozzi@tgen.org & lgarofano@tgen.org.